STOmics Standard Analysis (SAW)
The Stereo-seq Analysis Workflow (SAW) integrates multiple Stereo-seq spatial gene expression analysis tools. These tools can be used to reconstruct and visualize the spatial expression information of sequencing data on the chip. After analyzing the raw Stereo-seq sequencing data with SAW, a spatial expression matrix is obtained, which can be used for downstream analysis.
1.Main Function
(1) Image Preprocessing
Based on track lines, microscope thumbnail files are stitched together, and preliminary tissue segmentation and cell segmentation are also performed.
(2) Data Filtering
Low-quality sequences, sequences that cannot be aligned to CID sequences, sequences that cannot be aligned to exon regions, multi-aligned sequences, and duplicate sequences are all strictly filtered and screened.
(3) Generating Expression Matrix
After processing the sequenced data through computational algorithms, gene expression matrices are generated based on alignment information. Additionally, expression matrices are also generated for different bins.
(4) Image Registration and Tissue Segmentation
Segmenting gene expression data in non-tissue-covered areas to remove background noise. Through image registration and tissue segmentation, the expression signals in tissue regions can be accurately preserved.
(5) Spatial Clustering
This module performs unsupervised clustering of cells based on the expression matrix obtained from tissue segmentation, showing the spatial distribution of different cell types.
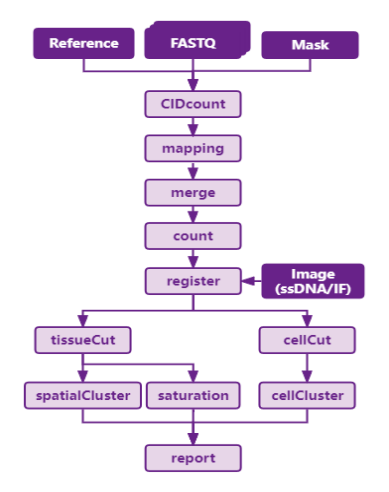
SAW Procedure
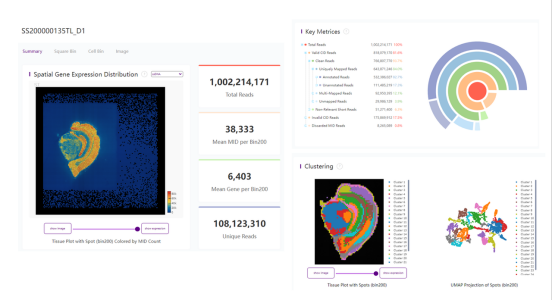
SAW Generated Analysis Report