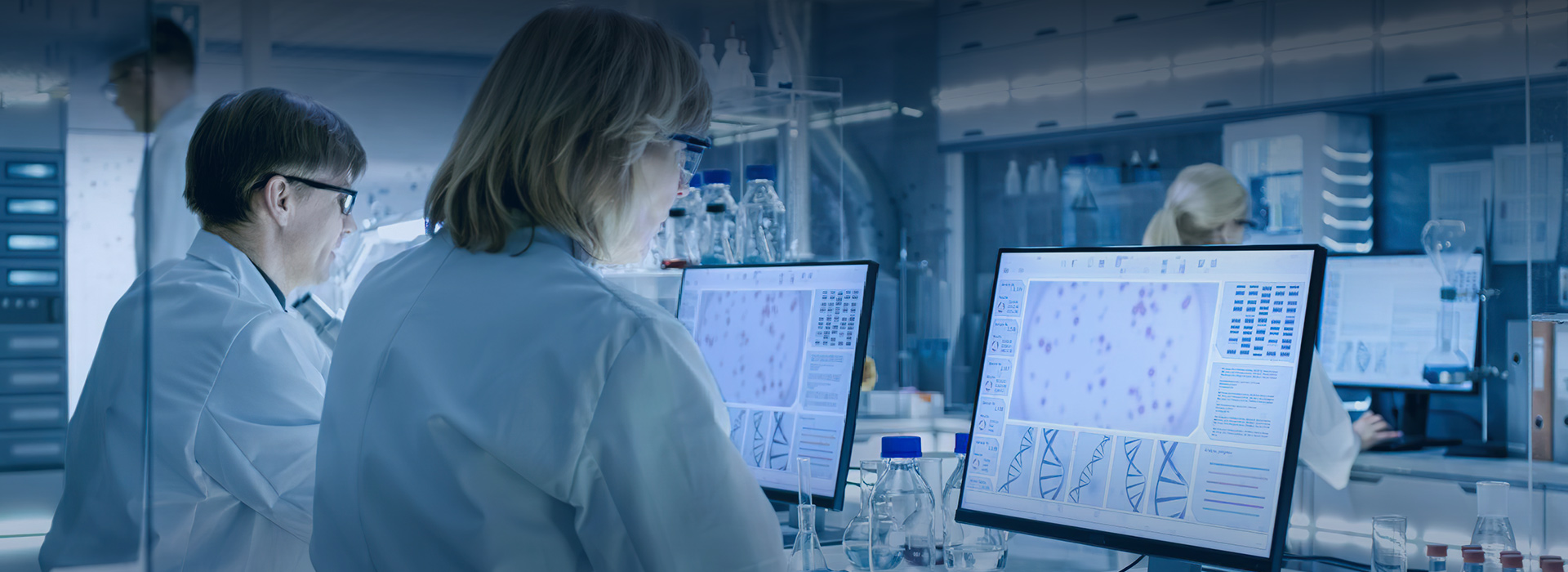
Data Mining supported by Stereopy
Stereopy is an open-source Python toolkit developed by BGI Research Institute, which can be used for data mining and data visualization in spatial transcriptomics. Stereopy provides a range of analytical functions for STOmics data, including cell clustering, spatial gene expression patterns, image segmentation, and more, to meet various needs. Meanwhile, performance and computational efficiency continue to be worked on improving.
Main Function
1. More suitable for downstream analysis of Stereo-seq spatial transcriptomics data.
2. Efficient reading, writing (I/O), preprocessing, and standardization of multiple spatial transcriptomics data formats are supported.
3. Developed Gaussian smoothing models, tissue and cell segmentation algorithm models, and cell correction algorithms.
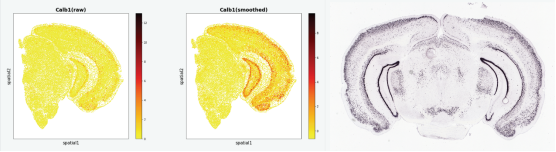
A heatmap of the coronal section of the mouse brain from the Allen Brain Atlas, generated using the Gaussian smoothing model in Stereopy.
Retrieved from https://mouse.brain-map.org/-experiment/show/71587929
4. Integration of various functionalities including dimensionality reduction, spatial clustering, cell clustering, and spatial expression pattern analysis.
5. Interactive visualization capabilities have been developed based on the characteristics of the Stereo-seq workflow.
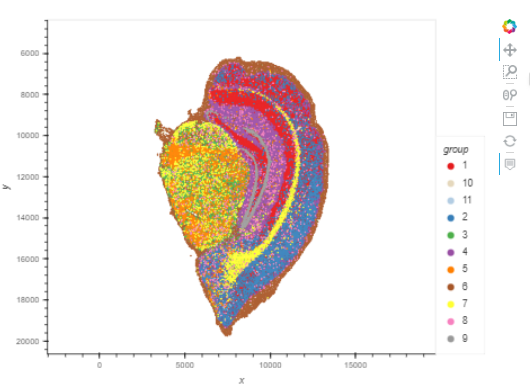
The Stereopy interactive visualization interface
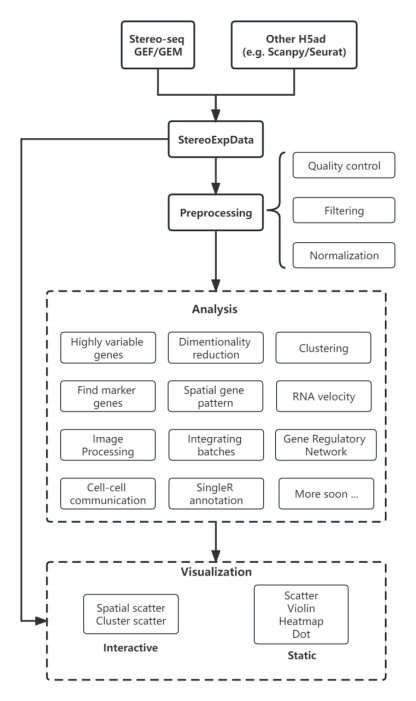
Stereopy流程图
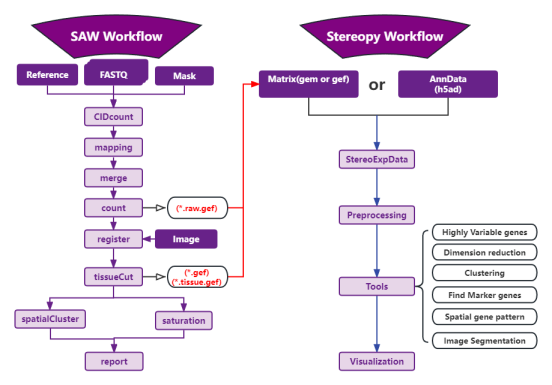
Workflow Chart